By Dishani Sen
Digital trace data refers to the electronic data that is generated as people interact with digital platforms, such as social media, online search engines, and mobile applications. These data have become valuable sources of information for businesses and researchers, particularly in the field of public health. The everyday opinions expressed on social media provide promising opportunities for measuring population-level statistics for health metrics; once such being disease prevalence. In this blog, we will explore how the same digital trace data can be utilised to predict the spread of Covid 19 infections across municipalities in Belgium.
Introduction
The outbreak of 2019’s disease coronavirus (COVID-19) is one of the worst recorded in history. According to the Institute for Health Metrics and Evaluation (IHME) at the University of Washington, the true global death toll is more than double the reported figures. Countless people that die while contaminated with SARS-CoV-2 are never tested for it, so their counts are not included in the official totals. Therefore, it is nearly impossible to investigate all the symptoms of the infection by relying only on health records.There are more people who have been infected with COVID-19 virus than it is ever recorded. To better understand the full spectrum of prevalence of COVID-19, the symptoms experienced by infected people and make further inferences regarding the spread of the infection among people, there is a need to look beyond hospital- or clinic-focused studies. Therefore, researchers have been looking at how digital trace data can be used to predict the spread of the virus in different communities. In this blog, such an alternate way is proposed from digital-trace data. It is a particularly innovative approach using publicly available social media data to track the spread of the disease in real time. The key problem addressed in this blog is predicting disease prevalence from signals in social-media data, especially Twitter.
Is it possible to predict the area-level prevalence of COVID-19 infections in Belgium by analysing self-reported symptoms on Twitter?

With 436 million monthly active users in 2022, Twitter was one of the most popular social media sites in the world. Twitter has long been a large source of communication for people all over the world, but it also contains a wealth of information that has grown increasingly vital for businesses and researchers to obtain valuable insights into trends, consumer behaviour, and public sentiments.
In this blog, a complete end to end pipeline to perform real time predictions of disease prevalence at a granular level in a population using social media (Twitter) data is presented. This proof of concept is performed for Covid-19 infections in Belgian municipalities. The results are highly encouraging, as estimates correlate with true disease prevalence for a selected time-frame with a coefficient greater than 0.9, showing promise for real-time disease prevention and monitoring.
Problem Statement:
Social media data have a potential application in the early identification of novel virus symptoms in digital epidemiology. There’s an optimistic future in the possibility that monitoring social media data as a viable strategy to public health surveillance. It is a critical competency that public health organisations are investing into in order to receive real-time signals of pandemic upticks and spread. However, social media data is often unorganised, and a non-representative sample of the population due to demographic skew in usage frequencies and access rates. As such, any direct estimate from a platform like Twitter is likely biased towards certain demographics. With this in mind, an attempt is made to use tweets (digital trace data) to make inferences about the granular level prevalence of COVID-19 infections in Belgium.
The goal of the POC presented here was to determine how this digital trace data, which is unstructured, non-representative, and biased, might be used to make inferences about the granular level prevalence of COVID-19 infections in Belgium.
Approach:
In this POC, a complete pipeline is constructed that includes collecting social media data, organising it into survey-like objects, and finally fitting a mixed effects multi-level regression model with post-stratification (MrP) to forecast COVID-19 cases on a specific day at the municipality level in Belgium.
Through this POC, the following contributions are made to three core elements : collecting mass scaled tweets, extracting demographic features and assigning a location value to convert unstructured digital data to survey-like objects, and using a multi-level regression model with post-stratification to make real-time predictions on the population using digital trace data.
The methodology can be broken down into the following parts:
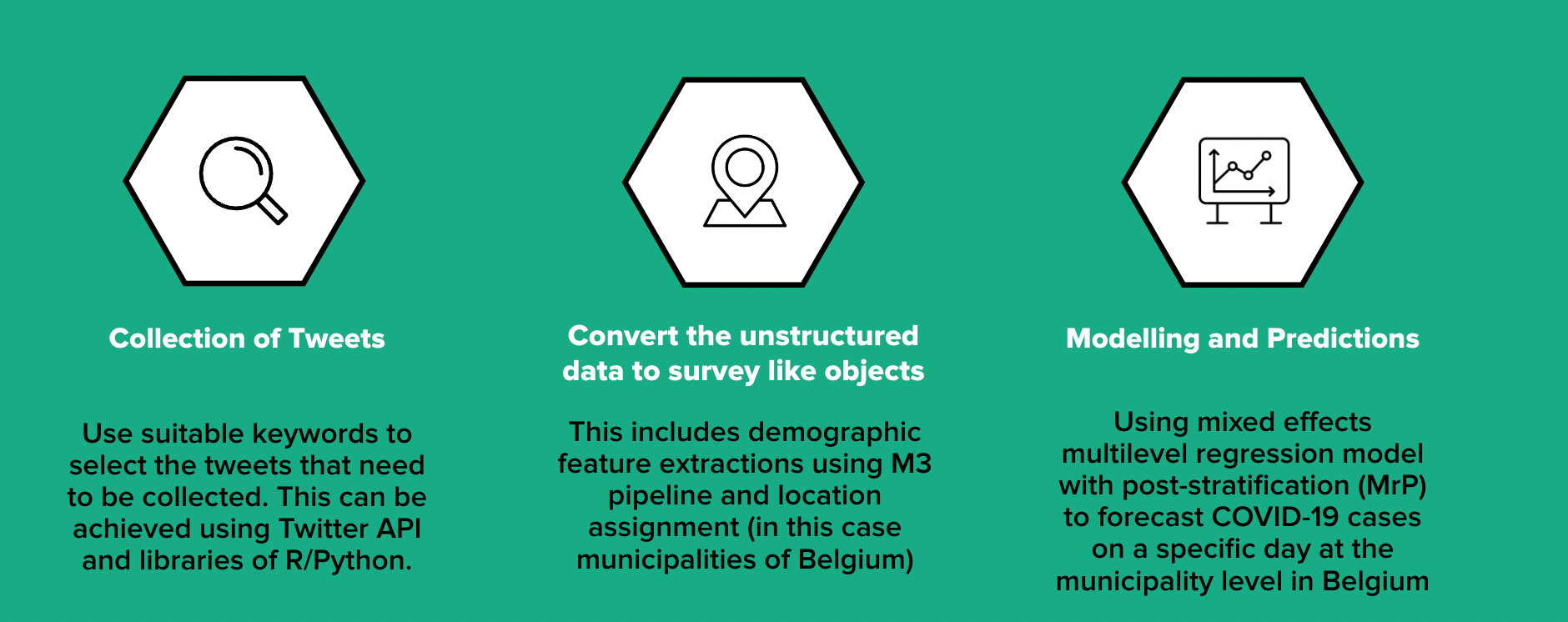
1) How is the mass scaled digital trace data (tweets) collected?
The methodology involves collecting tweets written in English, French and Dutch between March 1, 2020 and February 29, 2022 based on certain keywords related to symptoms of COVID-19 using Twitter API and R library "academictwitteR".
The keywords are a combination of COVID-19 synonyms (like coronavirus, corona, etc ) and common COVID-19 symptoms recommended by the CDC. To account for the second and third wave of the pandemic, the keywords are inclusive of the two later variants of COVID-19, the delta variant and the omicron variant. Tweets including important COVID-19 phrases along with phrases for the different variants of the virus (e.g., corona, covid, delta and omicron) as well as at least one of the symptoms are collected. Thus, tweets that have been collected using the infection-related keyword like "corona fever" and these self-reported symptoms are treated as “cases” where there is evidence of its corresponding user being infected by the COVID-19 virus.
An example of tweet that have been collected:
"I got a Covid test today because of a slight fever. I think it will be something else, which I have an appointment for tomorrow but it still feels weird. I prefer working from home with the kids to being ill with them by a long stretch. I can’t postpone being ill until they sleep.”
2) How is the unstructured digital trace data transformed into structured survey like objects amenable to statistical analysis?
To make the primary digital trace data amenable to further statistical analysis like post-stratification, important features like location, age and gender are prerequisite. Thus, this step is to extract those features and convert the cases into structured survey-like objects. The resultant data is a replication of results from a survey, containing, the age, gender, location and the date on which the particular individual has experienced symptoms.
2.1) How can we assign a location value to tweets for a granular level analysis?
In the context of this study, the location of interest is municipalities of Belgium.The “academictwitteR” package on R provides great flexibility in terms of control over the location from which tweets can be retrieved. The “point radius” argument of “academictwitteR” package can be utilised to regulate collection of data from every municipality of Belgium. Every municipality is conceptualised to be a circle in shape and tweets have been accumulated from every municipality, using the “point radius” argument, which takes, the latitude, longitude and radius as inputs. Information about the latitude, longitude and radius corresponds to the center point of each municipality. The R libraries, “BelgiumMaps.StatBel” which is contributed by StatBel Belgium and “sf” have been utilised to find the centers of each municipality in Belgium. The radius for each municipality is calculated by the formula radius= sq root (area of each municipality/pi) units.
2.2) How can we extract demographic features of the users which we know are important for the outcomes for COVID-19 pandemic (e.g., Age, Gender)?
M3 inference pipeline has been used on Python 3.6.6 via the package “m3inference” to gather the demographic information. M3 inference pipeline is a multilingual, multimodal, multi-attribute deep learning system for inferring demographics of users has been utilised to extract demographics information like age and gender of the cases. M3 inference pipeline is a deep learning system trained on a big Twitter data-set for demographic estimation. It has three primary characteristics: first, it is multimodal which means it accepts both visual and text input. The inputs are a profile image, a name (e.g., a natural language first and last name), a user-name (e.g., the Twitter screen name), and a brief self-descriptive text (e.g., a Twitter biography). Secondly, M3 is multilingual, operating in 32 main languages. And finally, M3 is multi-attribute which means that the model can predict three demographic attributes at the same time (gender, age, and human -vs - organisation status).The output from M3 inference is in probabilities (with 0 to 1) corresponding to four classes of age; age less than equal to 18, age 19-29, age 30-39 and age more than equal to 40, two classes of gender; male and female, and two classes of organisation; non-organisation and is organisation. The class with the highest probability for each attribute is the predicted class.
On a large-scale analysis of Europe, M3 inference pipeline model can reliably infer regional population counts and give demographic adjustments to selection biases. Thus key demographic features of interest are extracted using the state of art M3 deep learning pipeline, effectively transforming the unstructured twitter sample into a survey-like object.
3) How can these biased survey-like objects be utilised for generating representative real-time estimates of COVID-19 cases at municipality level in Belgium?
The survey like objects are now structured and have key demographics like age, gender, and a geo-location associated to municipalities in Belgium, and are thus amenable to statistical analysis. But they are still non-representative.
The technique of choice and relevance for the next step is a multi-level regression model with post stratification. MrP is a prominent method for adjusting non-representative samples in order to analyse opinion and other survey results more effectively. It uses a regression model to link individual-level survey results to multiple factors before rebuilding the sample to better reflect the population. MrP can thus not only provide a better understanding of behaviours, but also enable to analyse data that would normally be illegible for statistical inference.
MrP as a statistical technique is utilised to produce estimates of disease prevalence for small defined geographic areas. In general, MrP is an excellent technique to achieve certain goals, but it is not without drawbacks. If there is a biased survey, then it’s a great place to start with MrP but it’s not, a miracle of course . In this case, the digital trace data are biased as it is not a probability sample. They are not an actual representation of the real population for a number of reasons: people on social media do not represent the actual population. Firstly, there’s a lack of representation of all age groups, social classes, on social media. And secondly, even if people use social media platforms, not all of them tweet about them getting infected and self-report their symptoms. Nevertheless, it is still attempted to make real time estimates about the disease prevalence of COVID-19 infections among Belgian population from digital trace data with MrP.
MrP fundamentally trains a model on the survey data and then applies that learned model to another data-set. There is a major benefit of using MrP, that it allows to 're-weight' in a way that keeps uncertainty in mind and isn’t majorly hampered by small samples. In this case, small samples are in the form of cases pertaining to each municipality. The survey- like objects are modelled on the basis of their demographic characteristics and what is known about their area (i.e., which municipality of Belgium they belong to). This is the 'multilevel regression' part. There are elements of a person’s life, like age, gender and the municipality of Belgium where he/she lives, and these may provide an indication as to their likelihood of being infected by the disease in a certain way. The MrP technique allows us to model, based on demographic characteristics, the disease prevalence. So, this technique will assess the likelihood of an individual with a particular set of variables to be infected. “Multi-level regression” examines to what extent each of these elements has an effect on having the infection. In the subsequent 'post-stratification' stage, the census data is used to calculate how many people of each demographic type live in each area and combine this with additional relevant contextual information to predict how many of these people will be infected.
Multilevel regression for MrP is often estimated using Bayesian approaches, and this is the decision in this POC study as well, hence Stan and the 'rstan' package on R are used to carry out the modelling. A mixed effect multi-level logistic regression model is fit where fixed effects are used for intercept, individual level regressors (age, gender) and area level regressors to leverage correlation across areas over certain variables like income and demographic profile, whereas random effects are used for area level identifiers, in this case a nested index for municipalities.
Therefore, based on these demographic features and census characteristics, a mixed effects logistic regression model with post-stratification according to the Belgian census is proposed to forecast the number of infected individuals on a particular day.
Results:
In the case of this POC, the expectation from the model is not to make accurate predictions but to show that there is at-least some correlation with the model estimates and the actual observed number of cases per municipalities. Through that, it is established that there is feasibility in the approach used in the study, that is, building a pipeline from scratch to collect, organise and analyse social media data to predict COVID-19 infection prevalence. The results can be refined by improving the model and in future similar pipelines can be put in production for real time estimation of infection.
The Pearson’s Correlation Coefficient between the predictions based on cases count of 23 January 2022 from the model for the next day and the actual number of cases reported on 24 January 2022 is 0.938. This strong positive correlation is a very promising indication that there is enormous signal in the twitter data. On the other hand, the RMSE is found to be very high at 16241.3. The predictions are massively inflated, because the artificial split of the cases and controls is 50:50 which has increased the baseline rate dramatically.
Discussions:
The approach discussed here has several advantages over traditional surveillance methods. Firstly, it is free for the academia (however, business and industry professional might have to incur fees for using Twitter API), and the data is easily accessible - there are no specific hardware or software requirements to use it (beyond the toolkit of a laptop, R, Python and Stan) and the method does not require any additional infrastructure. This serves as an inexpensive way to get hands on important data as compared to physical surveys. Secondly, it is in principle able to track the spread in real-time; meaning it is possible to monitor new cases as soon as they emerge. Finally, it is based on completely anonymised data of users online who have self-reported being infected by coronavirus and does not rely on physical self-reported information from the population. After the data processing step, i.e., turning Twitter data into survey like objects, the information becomes anonymous; there is no personally identifying information. The subjects of the data, whether infected or not, is no longer recognisable. These advantages make social media monitoring an attractive field of research for tracking epidemics and monitoring the effectiveness of public health interventions. It can also be used to identify areas at high risk of the disease as well as people with potentially serious symptoms who may require further medical attention.
This POC has been ambitious in its goal to be able to use social media data to generate representative real time predictions of COVID-19 infection cases at the municipality level in Belgium, and the pipeline presented has ample opportunities of improvement. However, this is a back-bone proof of concept study that tries to show that, it is indeed possible to use statistical techniques, to utilise the untapped resource of social media data into digital epidemiology and study the prevalence of any disease at a granular level. Even though there are certain un-ignorable pitfalls, such as high incidence of sampling bias and inefficient geo-coding of the collected data, it is believed that these shortcomings can be overcome with further advanced research in the area. The main strength of the approach is that it confirms that there is enormous signal in social media data. The possibility of using historical data from social media platforms to make predictions of possible new cases at a municipality level is established. This offers tremendous potential for real-time disease spread monitoring and control at a granular level, here the municipal level in Belgium.
Conclusions:
A multitude of research have been done as part of the reaction to this pandemic in an effort to improve our understanding and prevent the virus's spread. A very novel strategy has been revealed in this blog, namely the use of publicly available social media data to track the spread of the disease in real time. In conclusion, it is suggested that the end-to-end pipeline presented in this POC represents a very valuable source of information for the scientific community and public health authorities. This approach has the potential to make a significant contribution to the fight against any infectious disease, such as COVID-19, by accelerating the rate at which new cases are detected, using sources other than official data and carrying out informed targeted interventions to mitigate the spread of the disease.
The use of self-reported symptoms on Twitter to predict disease prevalence has several implications for public health. For example, this approach could be used to identify areas where there is a higher risk of COVID-19 infections. This information could then be used to inform public health interventions, such as targeted testing and contact tracing.
Next Steps
As an outlook for the future, it would be interesting to extend the scope of utilising digital trace data from only Twitter, to other social media platforms like Reddit, Facebook, etc. Reddit features a robust API that permits users to access most of the information on the site. Further, an online dashboard available publicly that will enable users to access results of real-time predictions of progression of disease outbreak and track symptoms at their own province or municipality level is another goal that can be achieved in the future. This would allow users to compare the results of their municipality with those of other municipalities as well as healthcare officials to better manage emergencies by proper resource management and planning to tackle any novel outbreak.
In a Nutshell
An end-to-end pipeline with multi-level regression and post-stratification modelling approach is attempted and possible in the context of Belgium to solve the challenge of obtaining representative estimates of disease prevalence at the area-level using social media data. The approach proposed in this blog can achieve the objective of precisely forecasting the number of COVID-19 infections for a particular day by using social media data turned into survey-like objects along with the previous day’s COVID-19 cases count. While there are limitations to this approach, it has the potential to be a valuable tool for public health authority.
The potential approach developed and the results of this thesis have won the European master thesis award in statistics in 2023.

You might also like



