By Murilo Cunha
If you're not new to Python and data science, you probably heard of Jupyter notebooks. But if you haven't, here's the gist: it's an interactive environment, meaning you can run little bits of code and see the output, store variables in memory, etc.
That makes notebooks a good tool for experimentation, reporting and visualizations. And because of that, it's a popular tool of choice for data science in general. And this is why you see a lot of notebooks in places like Kaggle.
Personally I think notebooks are a good tool for figuring out what you want (think of experiments and proofs-of-concept), and once you do, I'd move everything to scripts and modules (think production code). But that's where the Python community gets divided. Because some people hate notebooks. Regardless of the stage of development you're in.
"I'd rather roll around in shattered glass naked than work with notebooks" - Someone, somewhere... probably
And honestly, I get it. As a machine learning engineer, I've had my pains with notebooks. That's because as great as they may be for quick prototyping and experimentation, as soon as you want to work with someone else, it gets equally as painful. Notebooks are filled with metadata (execution counts, kernel things, and other stuff that you just don't care about) that can cause git conflicts and leaving you to open a lot of nonsensical JSON files to resolve. As a quick example, just by opening someone else's notebook you will change some metadata, because notebooks include some kernel information (kernel specs and language to name a few).
Calls were heard, prayers answered. A family of tools emerged with the purpose of relieving those pains. And recently, it got a new family member: databooks.
databooks --help
databooks is a black-like tool for sharing and caring about notebooks. It has (currently) two functions: clearing metadata and resolving git conflicts.
databooks meta --help
Much like black, the only thing you need to pass is a path. We have sensible defaults to do the rest.
databooks meta path/to/notebooks"What are these defaults?", you ask? Great question. For each notebook in the path, by default:
- It will remove execution counts
- It won't remove cell outputs
- It will remove metadata from all cells (like cell tags or ids)
- It will remove all metadata from your notebook (including kernel information)
- It won't overwrite files for you
But the user is king, and as such you can still change all that. You could choose to remove cell outputs by passing --rm-outs. Or if there is some metadata you'd like to keep, such as cell tags, you can do so by passing --cell-meta-keep tags. Also, if you do want to save the clean notebook you can either pass a prefix (--prefix ...) or a suffix (--suffix ...) that will be added before writing the file, or you can simply overwrite the source file (--overwrite).
If you don't like typing every time you want to run the command, you can specify any argument values in the configuration file, which is a pyproject.toml in the root of your project. databooks finds this file by default, but you can also specify the config via --config path/to/your/config/pyproject.toml.
Make sure to check the docs for more info on the configuration and all the arguments that you can pass in the CLI.
databooks fix --help
As one would know, (git) conflicts are, unfortunately, part of life. No matter how much we try to avoid them. If in meta we try to avoid git conflicts, in fix we are fixing these conflicts after they have appeared. Similarly to databooks meta ..., the only required argument here is a path.
databooks fix path/to/notebooksWhat will it do? For each notebook in the path that has git conflicts:
- It will keep the metadata from the notebook in
HEAD - For the conflicting cells, it will wrap some special cells around the differences, like in normal git conflicts
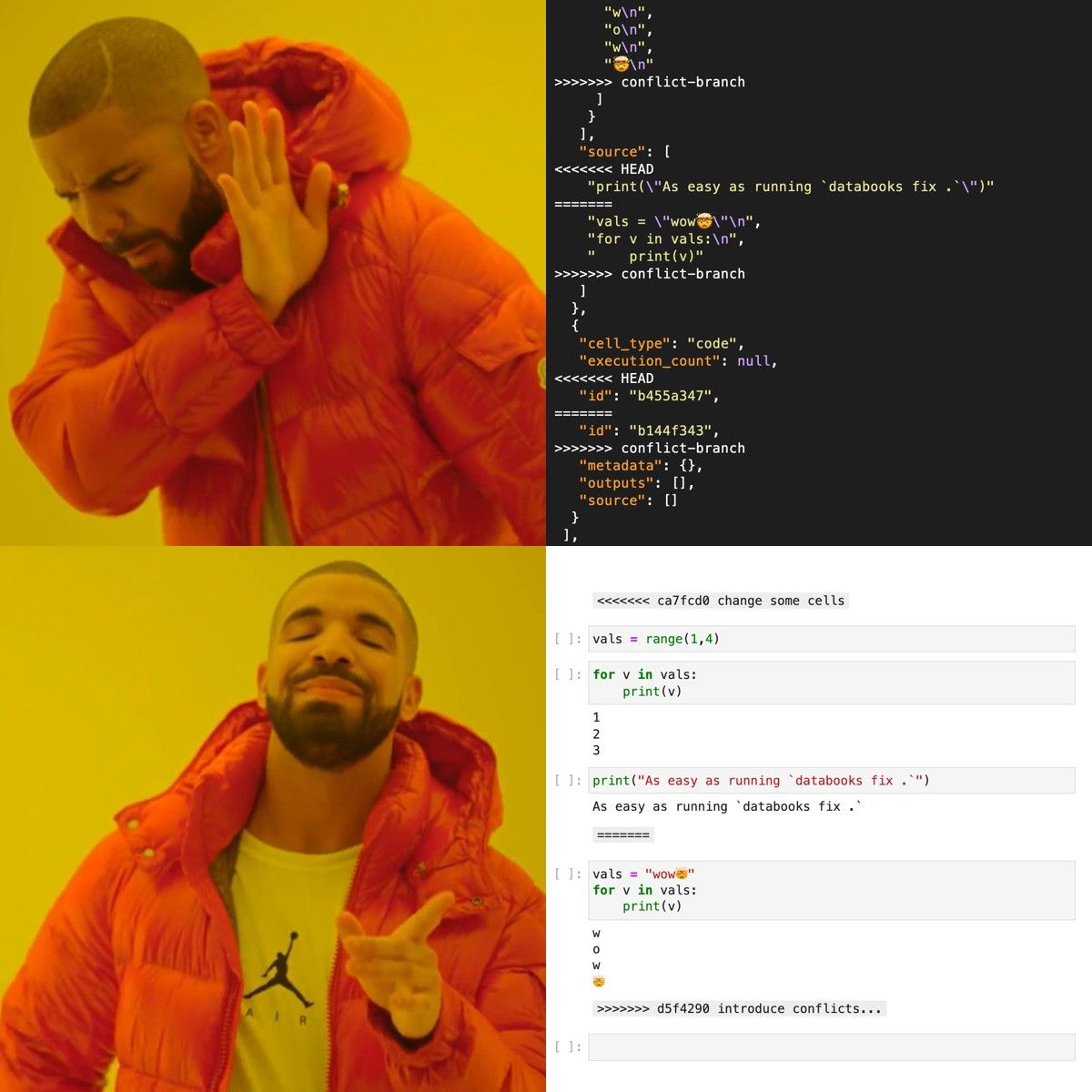
And as you could've guessed, you can still change the behavior. Similarly to what we saw above, this can be done with passing a configuration pyproject.toml file or specifying the CLI arguments. You could, for instance, keep the metadata from the notebook in BASE (as opposed to HEAD). If you know you only care about the notebook cells in HEAD or BASE, then you could pass --cells-head or --no-cells-head and not worry about all that.
You can also pass a special --cell-fields-ignore parameter, that will remove the cell metadata from both versions fo the conflicting notebook before comparing them. This is because depending on your Jupyter version you may have an id field, that will be unique for each cell. That is, all the cells will be considered different even if they have the same source and outputs as their ids are different. By removing id and execution_count (we'll do this by default) we only compare the actual code and outputs to determine if the cells have changed or not.
There is one tiny catch that if you commit a conflicted notebook without fixing it, databooks will not consider this a conflicted file anymore (just like git wouldn't). And since you've read this far, I'll share a little secret 🤫: we don't really "fix" the conflict, but rather we retrieve the previous file versions and compare them.
Once again, make sure the check the docs for more details!
Automate it
Now, as nice as this all may be, we can still do better. Having the tools is one thing, but that does't mean we'll always remember to use them.

Luckily, this is not a new problem. We can enforce standards via CI/CD. Or even better, we could use pre-commit hooks.
With that in mind, databooks has some examples in the docs on how to integrate the tool into the normal workflow, using GitHub Actions and the pre-commit package. Feel free to check them out!
And of course, all relevant info can be found on the repo! Thanks for reading! 🚀

